Unsupervised#
Outlier detection#
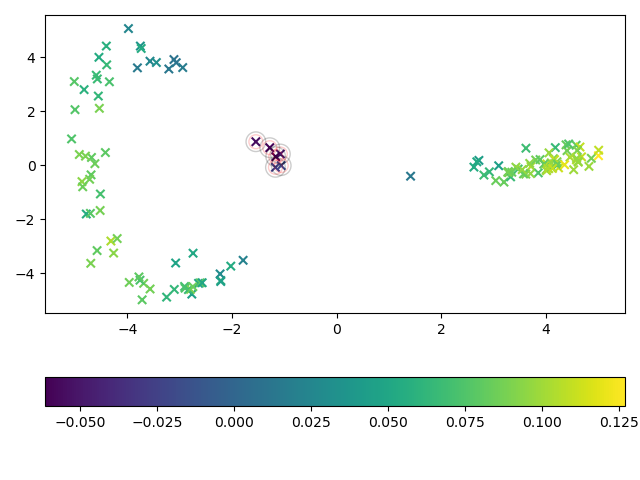
In this example, ShapeletIsolationForest is used to detect outliers. We
use the ShapeletForestEmbedding to visualize the time series and mark
each sample according to the outlier score assigned by the shapelet isolation forest.
True anomalous samples are encircled by a red circle and samples predicted as
anomalous are encircled by a black circle.
import matplotlib.pylab as plt
from sklearn.decomposition import PCA
from sklearn.model_selection import train_test_split
from sklearn.pipeline import make_pipeline
from wildboar.datasets import load_dataset
from wildboar.ensemble import ShapeletForestEmbedding, IsolationShapeletForest
random_state = 1234
x, y = load_dataset("CBF", repository="wildboar/outlier:easy")
x_train, x_test, y_train, y_test = train_test_split(
x, y, test_size=0.2, random_state=random_state, stratify=y
)
metric = "euclidean"
embedding = make_pipeline(
ShapeletForestEmbedding(
metric=metric, random_state=random_state, sparse_output=False
),
PCA(n_components=2, random_state=random_state),
)
isf = IsolationShapeletForest(
contamination=0.05,
metric=metric,
random_state=random_state,
n_jobs=-1,
)
isf.fit(x_train)
embedding.fit(x_train)
x_embedding = embedding.transform(x_test)
y_score = isf.decision_function(x_test)
y_pred = isf.predict(x)
fig, ax = plt.subplots()
mapping = ax.scatter(
x_embedding[:, 0], x_embedding[:, 1], c=y_score, cmap="viridis", marker="x"
)
ax.scatter(
x_embedding[y_test == -1, 0],
x_embedding[y_test == -1, 1],
edgecolors="red",
linewidths=1,
alpha=0.2,
facecolors="None",
s=100,
marker="o",
)
ax.scatter(
x_embedding[y_pred == -1, 0],
x_embedding[y_pred == -1, 1],
edgecolors="black",
linewidths=1,
alpha=0.2,
facecolors="None",
s=200,
marker="o",
)
plt.tight_layout()
fig.colorbar(mapping, ax=ax, orientation="horizontal")
plt.savefig("../fig/outlier_isf.png")