Explainability#
Counterfactual explanations#
wildboar can explain predictions of nearest neighbors classifiers and shapelet forest classifiers using counterfactual samples. In this scenario, counterfactuals are samples that are transformed such that the labeling of the sample changes. For instance, we might want to explain what changes are required to transforms a sample labeled as abnormal to normal. In this scenario, the normal sample would be the counterfactual sample.
In wildboar, counterfactual explainers are in the module wildboar.explain.counterfactual.
The easiest way to generate counterfactuals is to use the function counterfactuals:
from wildboar.explain.counterfactual import counterfactual
Note
Currently, the classifiers that supports counterfactual explanations
are ShapeletForestClassifier and KNearestNeighborsClassifier
from wildboar and scikit-learn respectively. Model agnostic counterfactual
explanations can be provided for other estimators.
To have more control over the generation of counterfactual samples, the classes
KNeighborsCounterfactual and ShapeletForestCounterfactuals can be used.
They implement the interface of BaseCounterfactuals which exposes two
methods fit(estimator) and transform(x, y), where the former fits
a counterfactual explainer to an estimator and the latter transform the i:th sample
of x to a sample labeled as the i:th label in y.
>>> from wildboar.explain.counterfactual import KNeighborsCounterfactuals
>>> from sklearn.neighbors import KNeighborsClassifier
>>> clf = KNeighborsClassifier(n_neighbors=5, metric='euclidean')
>>> clf.fit(x_train, y_train)
>>> c = KNeighborsCounterfactuals()
>>> c.fit(clf)
>>> counterfactual, success = c.transform(x_test, y_desired)
>>> counterfactual[success] # only successful transformations
Warning
KNeighborsCounterfactuals only supports KNeighborsClassifier fit
with the Euclidean distance.
Example#
In the following example, we explain the a nearest neighbors classifier and a shapelet forest classifier for the datasets TwoLeadECG and explaining samples classified as 2.0 if they instead where classified as 1.0 (in the legend denoted as abnormal and normal respectively).
import numpy as np
from sklearn.neighbors import KNeighborsClassifier
from sklearn.model_selection import train_test_split
from wildboar.datasets import load_two_lead_ecg
from wildboar.ensemble import ShapeletForestClassifier
from wildboar.explain.counterfactual import counterfactuals
x, y = load_two_lead_ecg()
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size=0.1, random_state=123)
# Change estimator to `KNeighborsClassifier(metric='euclidean')`
# to generate k-nearest counterfactuals instead
estimator = ShapeletForestClassifier(random_state=123, n_jobs=-1, metric="euclidean")
# fit the estimator to the training data
estimator.fit(x_train, y_train)
# only consider samples predicted as 2
x_test = x_test[y_test == 2.0]
# generate counterfactuals for the samples classified as 2, instead labeled as 1
x_counterfactuals, success, score = counterfactuals(estimator, x_test, 1.0, scoring="euclidean", random_state=123)
# only consider the successful counterfactuals
x_test = x_test[success]
x_counterfactuals = x_counterfactuals[success]
i = np.argsort(score[success])[:2]
x_counterfactuals = x_counterfactuals[i, :]
x_test = x_test[i, :]
Plotting the first two counterfactual samples with the lowest score yields the following
figure for the KNeighborsClassifier:
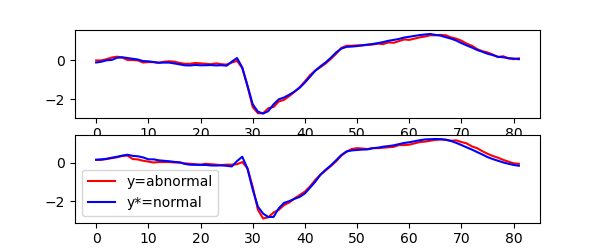
and for the ShapeletForestClassifier:
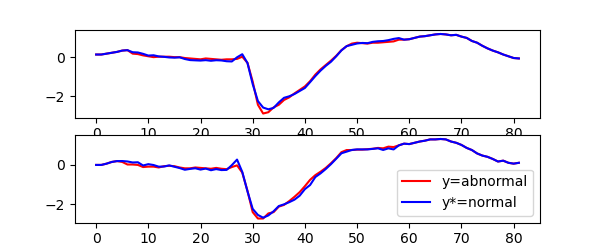
We can observe that the counterfactual explainer for nearest neighbors classifier tend to change larger parts of the time series, while the shapelet forest counterfactuals tend to have fewer and smaller changes.