Shapelets#
In this example, we will use random shapelets to transform time series.
[1]:
import matplotlib.pyplot as plt
import numpy as np
from sklearn.decomposition import PCA
from sklearn.pipeline import make_pipeline
from wildboar.datasets import load_dataset
from wildboar.transform import RandomShapeletTransform
from wildboar.ensemble import ShapeletForestEmbedding
random_state = 1234
First, we load the dataset.
[2]:
x, y = load_dataset("CBF")
labels, index = np.unique(y, return_inverse=True)
We define a function for plotting the 3d-projection.
[3]:
def plot(p, var, labels, index):
colors = plt.cm.rainbow(np.linspace(0, 1, len(labels)))
fig = plt.figure(figsize=(8, 8))
ax = fig.add_subplot(projection="3d")
ax.scatter(p[:, 0], p[:, 1], p[:, 2], color=colors[index, :])
ax.set_xlabel("Component 1 (%.2f variance explained)" % var[0])
ax.set_ylabel("Component 2 (%.2f variance explained)" % var[1])
ax.set_zlabel("Component 3 (%.2f variance explained)" % var[2])
Random shapelet embedding#
Next, we define the pipeline.
[4]:
rse = make_pipeline(
RandomShapeletTransform(
n_shapelets=1000,
metric="dtw",
metric_params={"r": 0.5},
random_state=random_state,
n_jobs=-1,
),
PCA(n_components=3, random_state=random_state),
)
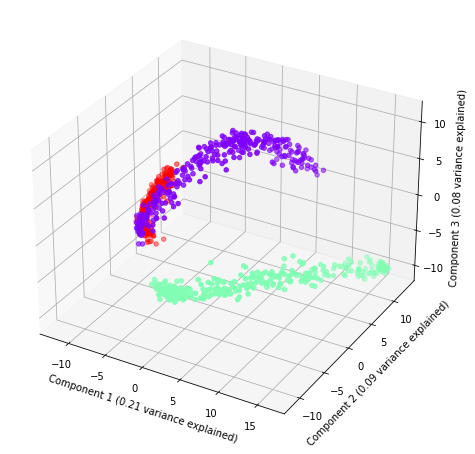
And, plot the three components with the amount of explained variance.
[5]:
plot(rse.fit_transform(x), rse.steps[1][1].explained_variance_ratio_, labels, index)
Shapelet forest embedding#
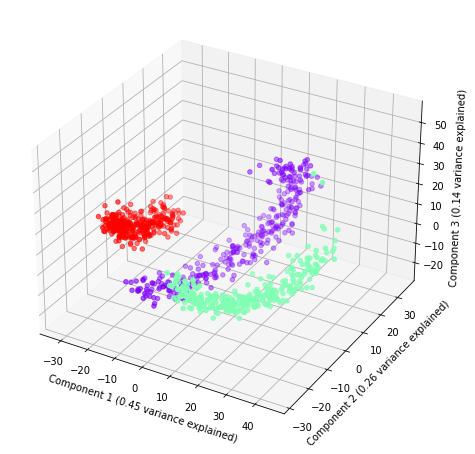
Finally, we define a new pipeline with ShapeletForestEmbedding and, again, plot it.
[6]:
sfe = make_pipeline(
ShapeletForestEmbedding(
n_estimators=1000,
metric="scaled_euclidean",
random_state=random_state,
n_jobs=-1,
sparse_output=False,
),
PCA(n_components=3, random_state=random_state),
)
plot(sfe.fit_transform(x), sfe.steps[1][1].explained_variance_ratio_, labels, index)